Agent Guide#
Use this page after OntoAnno is installed, configs/demo.yaml is configured,
and the web app opens correctly.
If you have not done those steps yet, start here first:
Panel Introduction#
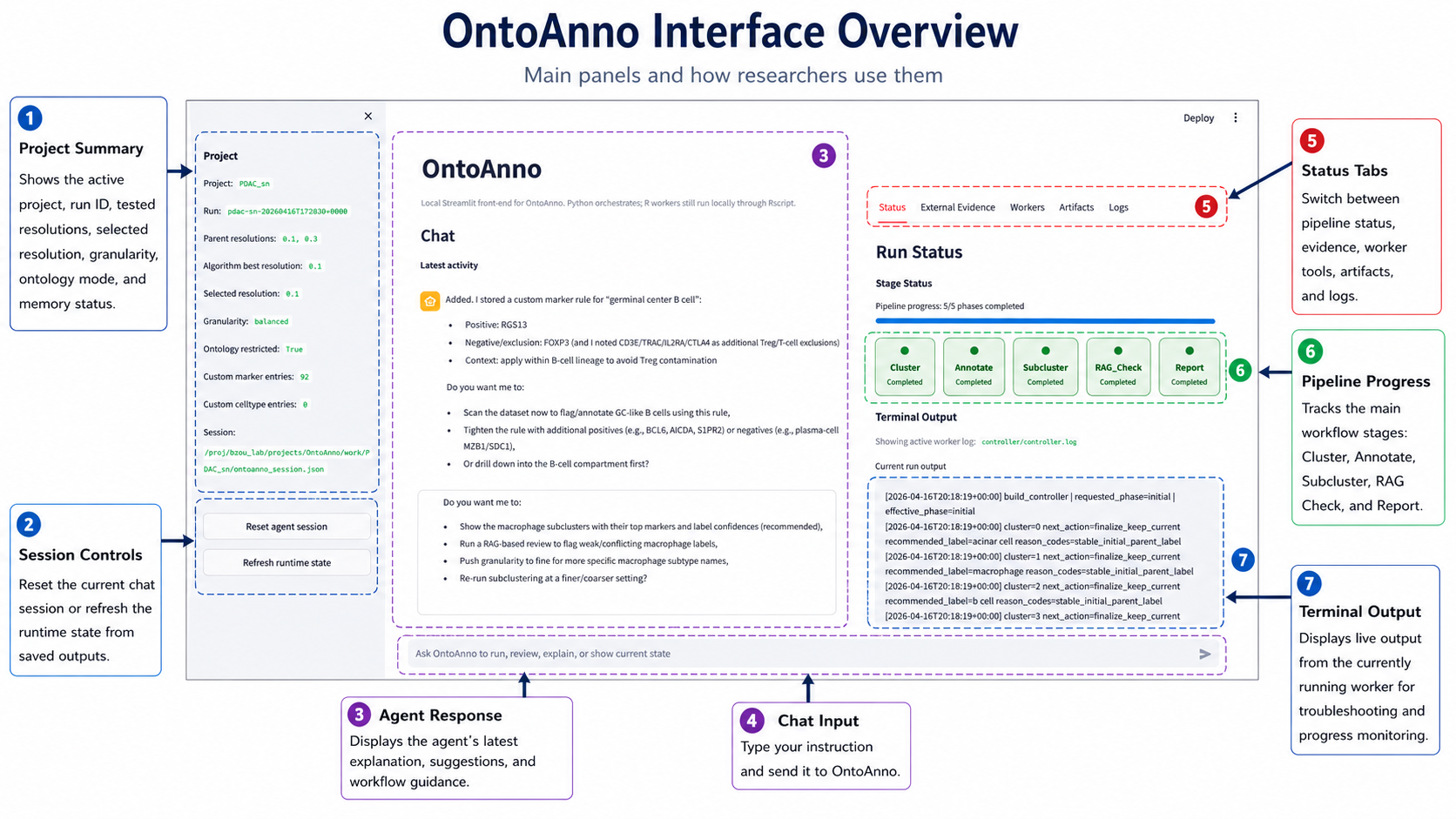
The OntoAnno page has three main working areas.
No. |
Panel |
What it shows |
When to use it |
|---|---|---|---|
1 |
Project panel |
Project name, run ID, parent resolutions, selected resolution, granularity, ontology setting, and evidence counts. |
Check this before running a workflow. |
2 |
Session controls |
Buttons for resetting the agent session or reloading the saved session from disk. |
Use only when you want to clear the chat or refresh stale page state. |
3 |
Agent response |
Agent replies, reasoning summaries, completed actions, suggestions, and follow-up questions. |
Read this after each request to confirm what OntoAnno did or what it needs next. |
4 |
Chat input |
Natural-language box for running annotation, review, evidence, subcluster, and report tasks. |
Type what you want OntoAnno to do. |
5 |
Status tabs |
Tabs for pipeline status, external evidence, manual workers, artifacts, and logs. |
Switch panels when monitoring runs, reviewing evidence, or inspecting outputs. |
6 |
Pipeline progress |
The main workflow stages: Cluster, Annotate, Subcluster, RAG_Check, and Report. |
Watch this to see which stage has completed or is currently running. |
7 |
Terminal output |
Live output from the currently running worker. |
Use this for progress monitoring and troubleshooting failed workers. |
Cluster#
The cluster module prepares cluster-level marker genes for parent annotation.
It uses the parent_res values from configs/demo.yaml.
If annotation.preprocess is true, OntoAnno first runs Seurat
normalization, variable feature selection, scaling, PCA, and UMAP. It then
clusters cells at each parent resolution and finds marker genes for each
cluster.
Typical chat request:
Run the parent annotation
This request starts the parent workflow, so the cluster module runs first when cluster outputs are missing.
Check results in:
Statusfor cluster progress and logs.Artifactsfor resolution-specific marker and prediction outputs.
Annotation#
The annotation module assigns parent cell-type labels to clusters. It uses cluster marker genes, tissue/species context, ontology settings, and repeated LLM annotation runs.
Typical chat request:
Run the parent annotation
After it finishes, ask:
What is the selected resolution?
Check results in:
Artifactsfor the parent annotation table.Artifactsfor the prediction figure.Artifactsfor the resolution score table.
If the selected resolution is not what you want, tell the agent directly:
Change the resolution to 0.3
Changing resolution changes which cluster assignment is used. It is different from granularity, which is handled during RAG-based review and label refinement.
Subcluster#
The subcluster module looks deeper into one parent cell type. Use it when a major parent label is biologically broad or contains multiple expected subtypes.
Typical chat request:
Look deeper into macrophages
OntoAnno subsets that parent cell type, reclusters it using sub_res, and
generates subtype annotations.
Check results in:
Statusfor subcluster progress and logs.Artifactsfor subcluster annotation tables and figures.
Subclustering is optional. You can skip it and continue directly to RAG review or report generation.
External Evidence#
External evidence is project memory that OntoAnno can use later when reviewing parent or subcluster annotations. You can add it before or after annotation; if you add it early, OntoAnno stores it and applies it when the RAG review and LLM judge steps run.
To add marker evidence manually, say the cell type and markers explicitly:
Add these markers to pericyte: RGS5, CSPG4, MCAM
User-provided marker evidence is treated as high-priority evidence during review. It can support or challenge parent and subcluster labels during RAG review, but it does not change clustering, marker-gene detection, or the initial GPTAnno parent annotation directly. Only add it when the cell type and marker relationship is explicit.
You can also add literature-derived evidence from a PDF. Open External
Evidence and use the PDF extraction panel:
Upload a literature PDF.
Choose the page range for figure extraction.
Keep the default render DPI unless the figure text is too small.
Click
Extract PDF evidence.
OntoAnno extracts text evidence with the GPTAnno/PDF2markers text pipeline and extracts figure evidence with a vision LLM. The resulting celltype-marker pairs appear under literature-provided evidence. These PDF-derived entries are supportive evidence for review, not golden rules.
RAG_Check#
The RAG check module reviews annotation consistency using ontology candidates, reference marker evidence, user-provided evidence, literature-provided evidence, and LLM comparison.
Granularity belongs to this review stage. It controls how specific the reviewed label candidates should be after clusters and marker genes are already fixed. It changes which ontology candidates are compared by the LLM judge; it does not force the LLM to choose a more specific or more general label. Accuracy remains the first priority, so the LLM can keep the current label or send the cluster to human review if the markers do not support the requested specificity.
The reviewed labels are too coarse. Make them more specific.
Run it after parent annotation or after subclustering:
Run the RAG check
Check results in:
Artifactsfor flagged clusters and candidate labels.Artifactsfor RAG marker evidence comparisons.External Evidencefor stored user and literature evidence.
If OntoAnno reports unresolved clusters, use the manual review controls in the RAG review panel. This is not a separate annotation algorithm; it is the final human resolution step for clusters that automated comparison did not safely finalize. You can also ask the agent to continue with manual review:
Continue with human review
For each unresolved cluster, choose the final label or enter a custom label. You can also save a specific decision through chat:
For cluster 3, label as cardiomyocyte
After the decisions are saved, ask the agent to export the reviewed labels:
Finish review and export the reviewed annotations
Reference Label Comparison#
If you have a CSV with known or manual labels, you can give it to the agent instead of editing optional YAML fields by hand:
Compare with /data/project/manual_labels.csv using column celltype
The agent will add the reference-label path and label-column name to the config and enable optional evaluation.
Report#
The report module creates the final report after annotation and review. The default output is HTML. A PDF report is also available when requested.
Typical chat request:
Generate the final report
To choose a specific format, say it directly:
Generate a PDF report
Generate an HTML report
The report summarizes parent annotations, selected resolution, RAG review, human review decisions, available subcluster results, figures, and output tables.
Check results in:
Artifactsfor the report preview.work/<project>/andruns/<run_id>/for saved output files.
Important Outputs#
OntoAnno keeps internal workflow files in both work/<project>/ and
runs/<run_id>/. To make the final outputs easier to find, important
user-facing files are also collected under:
work/<project>/results/
Large Seurat objects may appear there as symbolic links so that OntoAnno does
not duplicate large files. Open result_index.csv in that folder to see the
source path for every collected result.
File in |
What it contains |
When it appears |
|---|---|---|
|
Index of collected results, source paths, copy/link method, and descriptions. |
Whenever result files are synchronized. |
|
Selected parent resolution and cluster column. |
After parent annotation. |
|
Scores used to compare parent resolutions. |
After parent annotation. |
|
Seurat object with parent labels. |
After parent annotation. |
|
Per-cell metadata with reviewed parent labels. |
After RAG review is exported. |
|
Cluster-level final labels and decision sources. |
After RAG review is exported. |
|
Seurat object with reviewed parent labels. |
After RAG review is exported. |
|
Per-cell metadata with final subcluster labels. |
After subcluster annotation. |
|
Seurat object with final subcluster labels. |
After subcluster annotation. |
|
RAG check outcome for each cluster. |
After RAG check. |
|
LLM judge decisions and short reasons. |
After RAG check when LLM comparison was needed. |
|
Stored user-provided and literature-provided marker evidence. |
After external evidence is added. |
|
Final OntoAnno report. |
After report generation. |